The score is a number that reflects the conservation at a position.
position_aggregator: mean [default]
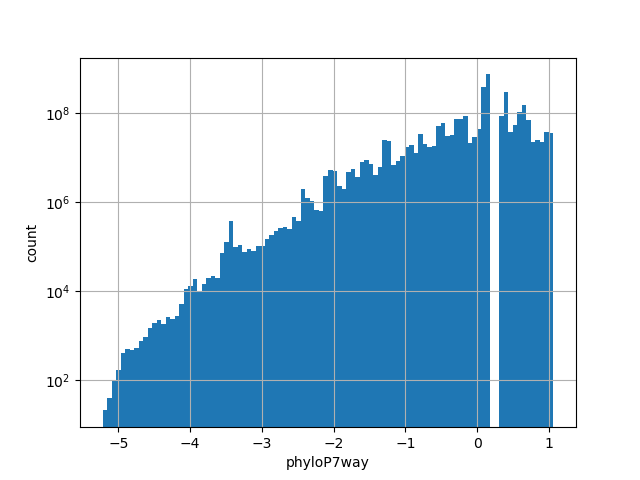
Annotator to use with genomic scores depending on genomic position like phastCons, phyloP, FitCons2, etc.
| Id | pipeline/Paper_pipeline |
|---|---|
| Type | annotation_pipeline |
| Version | 0 |
| Summary |
Annotation Pipeline used in the paper.
|
| Description |
This is the example pipeline used in the GAIn paper. It includes a variety of annotators, such as effect annotation, position scoring, allele scoring, gene scoring, gene set annotation, and liftover annotation. The pipeline is designed to demonstrate the capabilities of the GAIn framework for genomic annotation. |
| Labels |
| Summary | Demo pipeline |
|---|---|
| Description | Demonstrates a GAIn pipeline |
| Input reference genome | hg38/genomes/GRCh38-hg38 |
Worst effect across all transcripts.
List of all genes
Comma separated list of all affected genes.
Annotator to identify the effect of the variant on protein coding.
The score is a number that reflects the conservation at a position.
position_aggregator: mean [default]

Annotator to use with genomic scores depending on genomic position like phastCons, phyloP, FitCons2, etc.
Normalized allele.
Alternate allele frequency
position_aggregator: mean [default]
allele_aggregator: max [default]
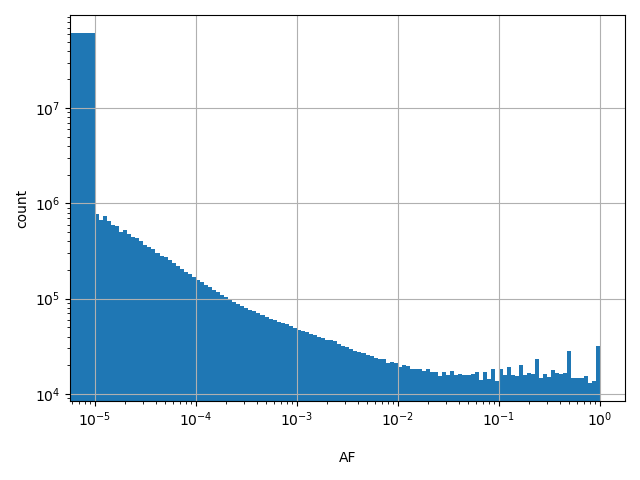
Annotator to use with scores that depend on allele like variant frequencies, etc.
normalized_alleleMVP scores were ranked among all MVP scores in dbNSFP. The rankscore is the ratio of the rank of the score over the total number of MVP scores in dbNSFP.
position_aggregator: mean [default]
allele_aggregator: max [default]
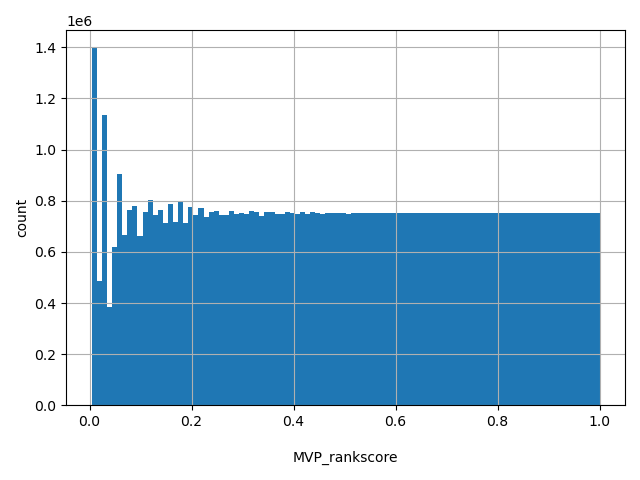
Annotator to use with scores that depend on allele like variant frequencies, etc.
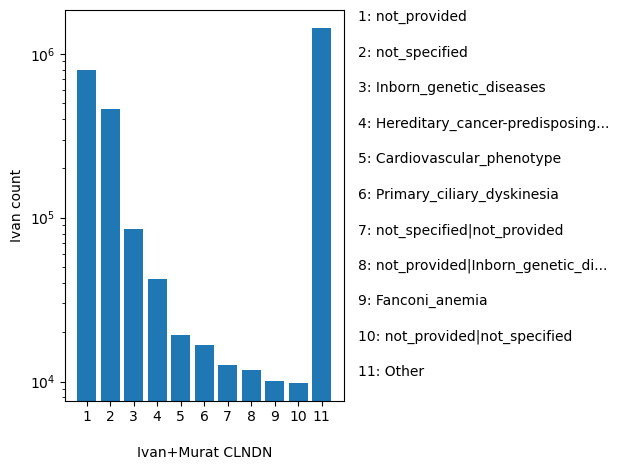
normalized_alleleAggregate germline classification for this single variant; multiple values are separated by a vertical bar
position_aggregator: join(,) [default]
allele_aggregator: join(,) [default]
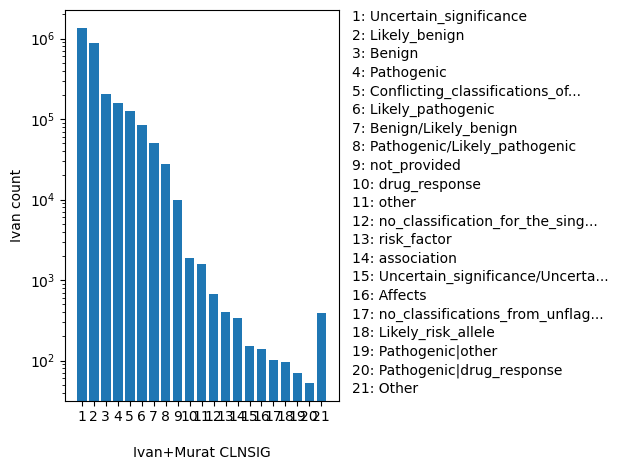
ClinVar's preferred disease name for the concept specified by disease identifiers in CLNDISDB
position_aggregator: join(,) [default]
allele_aggregator: join(,) [default]
Annotator to use with scores that depend on allele like variant frequencies, etc.
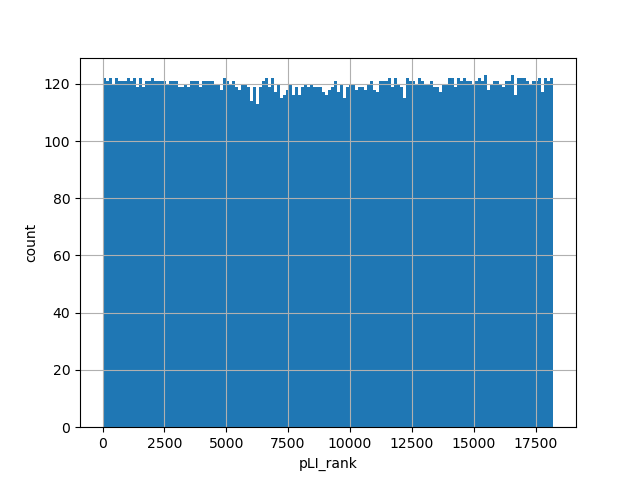
normalized_alleleGene rank after sorting by pLI intolerance score
gene_aggregator: dict [default]
replicationforkprocessing
This gene set collection annotator uses the GO gene set collection.
The lifted over annotatable
Annotator to lift over a variant from one reference genome to another.
Adipose Nuclei Score (E063)
position_aggregator: mean [default]
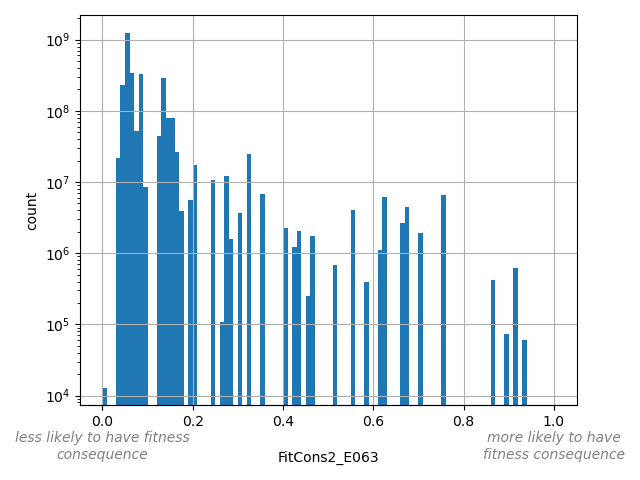
Annotator to use with genomic scores depending on genomic position like phastCons, phyloP, FitCons2, etc.
hg19_annotatable| Filename | Size | md5 |
|---|---|---|
| Paper_pipeline.yaml | 1.39 KB | dbb8df33c339024c42e7b589e529b22a |
| genomic_resource.yaml | 482.0 B | d746b1b4c60a7c2cf465dc2aed395630 |
| statistics/ |