phylop30way
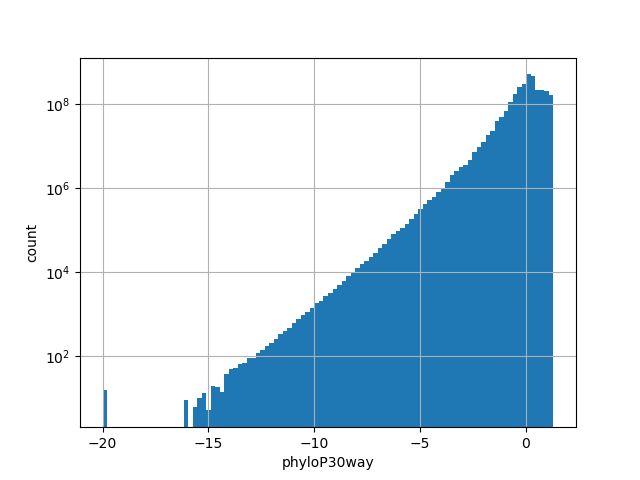
| Id | hg38/scores/phyloP30way |
|---|---|
| Type | position_score |
| Version | 0 |
| Summary |
Conservation score based on the multiple alignment of 30 species
|
| Description |
phyloP (phylogenetic P-values) scores measure the evolutionary rate at individual nucleotides or other genomic elements, providing a statistical framework to determine whether a particular site is evolving at a rate that is slower or faster than expected under a neutral model of evolution. phyloP30way refers to a specific set of phyloP conservation scores derived from a multiple sequence alignment of the genomes of 30 different species. This score is derived from a method called PHAST (Phylogenetic Analysis with Space/Time models), which uses multiple sequence alignments and phylogenetic trees to identify conserved elements across different species. Positive scores indicate conservation, meaning the site is evolving more slowly than expected under a neutral model (purifying selection). Negative scores indicate acceleration, meaning the site is evolving faster than expected (positive selection or fast evolution). PhyloP uses a likelihood ratio test to compare the likelihood of the observed data under a model of neutral evolution versus a model of constrained or accelerated evolution. The result is a P-value that reflects the statistical significance of the deviation from the neutral rate. Downloaded on: 06/10/24 https://hgdownload.soe.ucsc.edu/goldenPath/hg38/phyloP30way/hg38.phyloP30way.bw |
| Labels |
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| phyloP30way | float |
phylop30way |
The score is a number that reflects the conservation at a position.
Small values desc: less conserved
Large values desc: more conserved
|
 |
[-20, 1.31] |
| Filename | Size | md5 |
|---|---|---|
| genomic_resource.yaml | 2.21 KB | c8e8515c469e156860199832433ea1c3 |
| hg38.phyloP30way.bw | 7.82 GB | d68e11642735f4524fc519a1b6823cd0 |
| hg38.phyloP30way.bw.dvc | 105.0 B | 7de8282098f1c0d5562e1a1f1befc359 |
| statistics/ |