| ID |
Type |
Default annotation |
Description |
Histogram |
Range |
| deletion_duplication |
str |
deletion_duplication
|
duplication or deletion
|
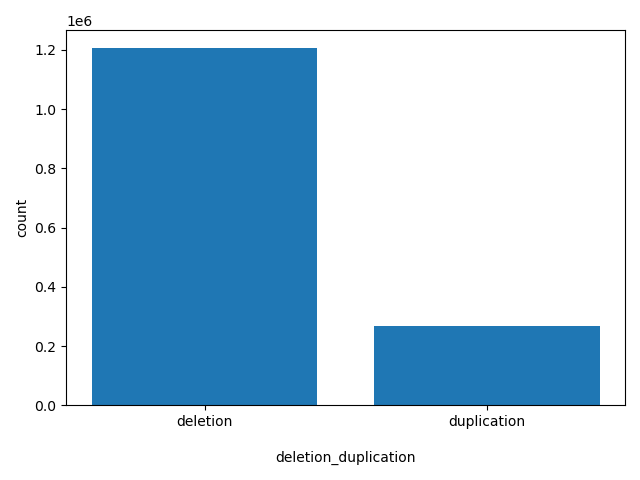 |
deletion, duplication |
| cnv_name |
str |
cnv_name
|
Handy name to refer to the CNV.
|
No histogram: Empty histogram for cnv_name in a region: Too many unique values 101 for categorical histogram. |
NO DOMAIN |
| SVLEN |
int |
SVLEN
|
SV length
|
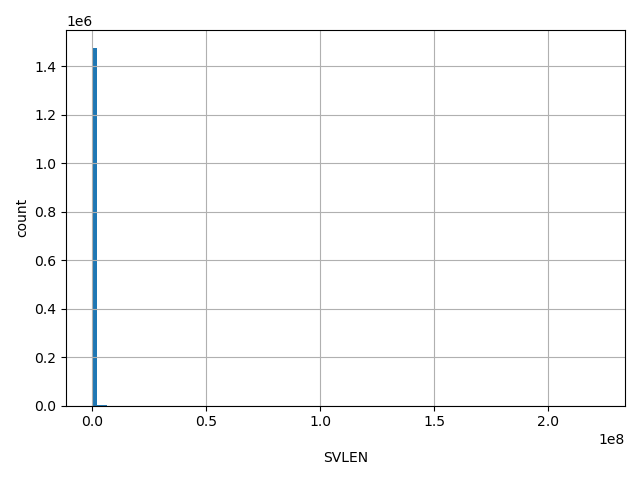 |
[50, 2.22e+08] |
| AN |
int |
AN
|
Total number of alleles genotyped (biallelic sites only).
|
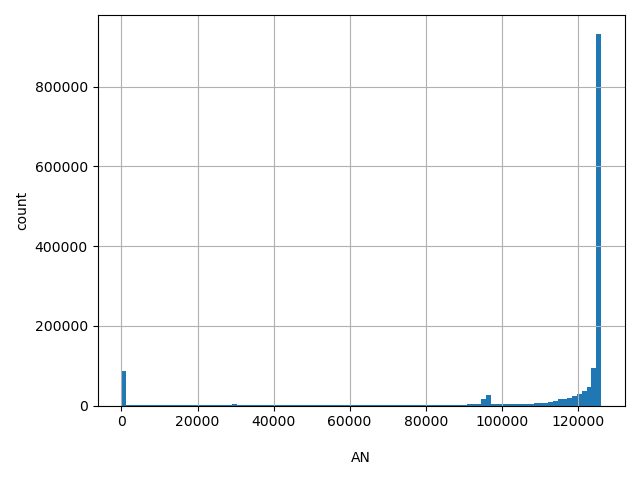 |
[1, 1.26e+05] |
| AC |
int |
AC
|
Number of non-reference alleles observed (biallelic sites only).
|
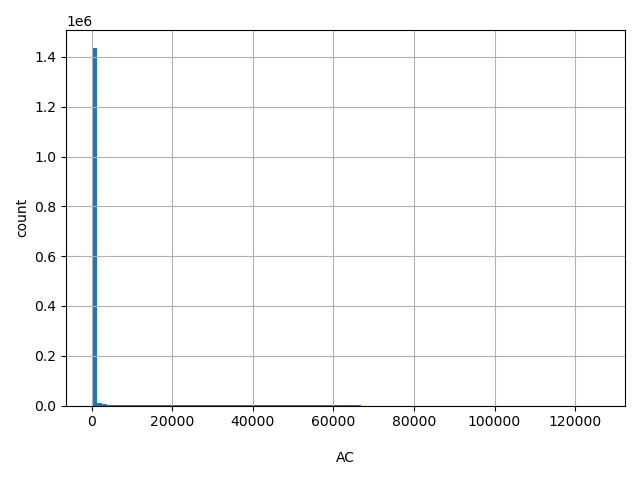 |
[1, 1.26e+05] |
| AF |
float |
AF
|
Allele frequency (biallelic sites only).
|
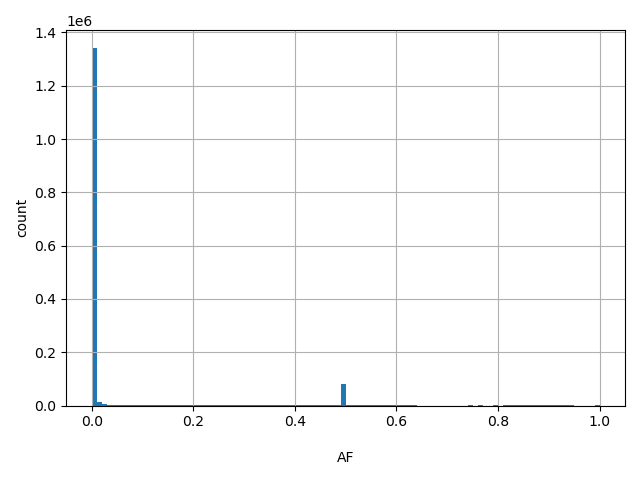 |
[8e-06, 1] |
| AN_afr |
int |
AN_afr
|
Total number of African-American alleles genotyped (biallelic sites only).
|
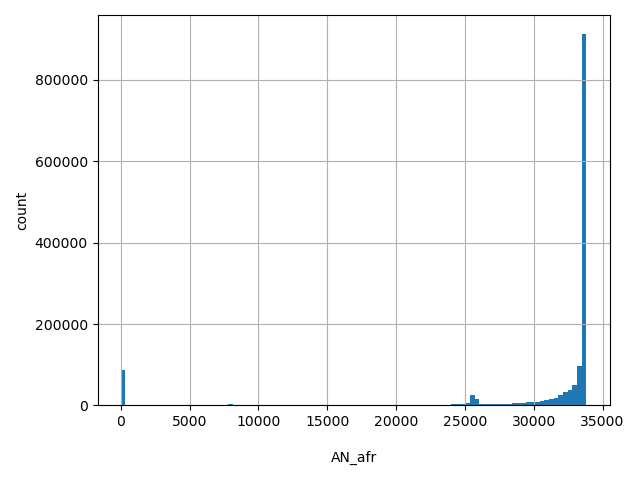 |
[0, 3.38e+04] |
| AN_amr |
int |
AN_amr
|
Total number of Latino alleles genotyped (biallelic sites only).
|
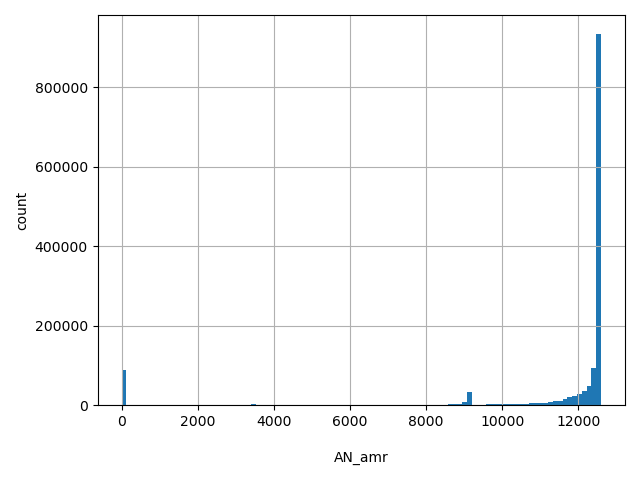 |
[0, 1.26e+04] |
| AN_asj |
int |
AN_asj
|
Total number of Ashkenazi Jewish alleles genotyped (biallelic sites only).
|
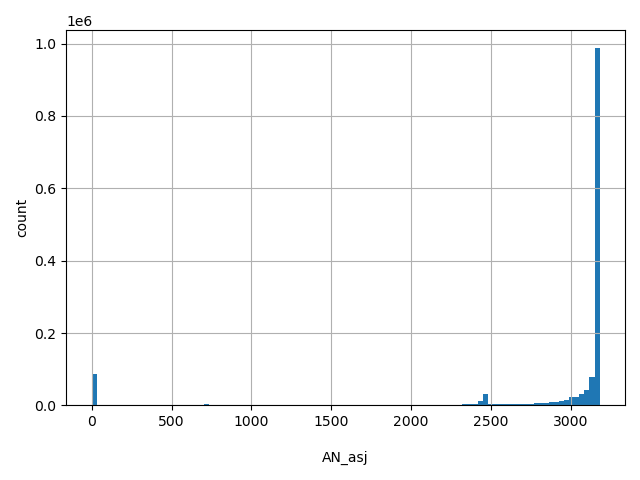 |
[0, 3.18e+03] |
| AN_eas |
int |
AN_eas
|
Total number of East Asian alleles genotyped (biallelic sites only).
|
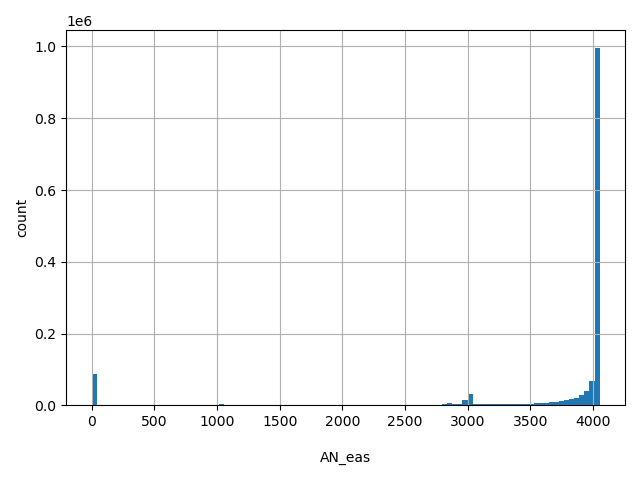 |
[0, 4.05e+03] |
| AN_fin |
int |
AN_fin
|
Total number of Finnish alleles genotyped (biallelic sites only).
|
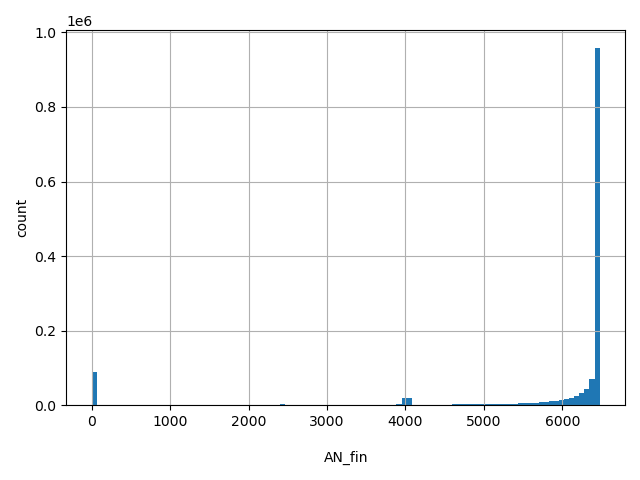 |
[0, 6.48e+03] |
| AN_mid |
int |
AN_mid
|
Total number of Middle Eastern alleles genotyped (biallelic sites only).
|
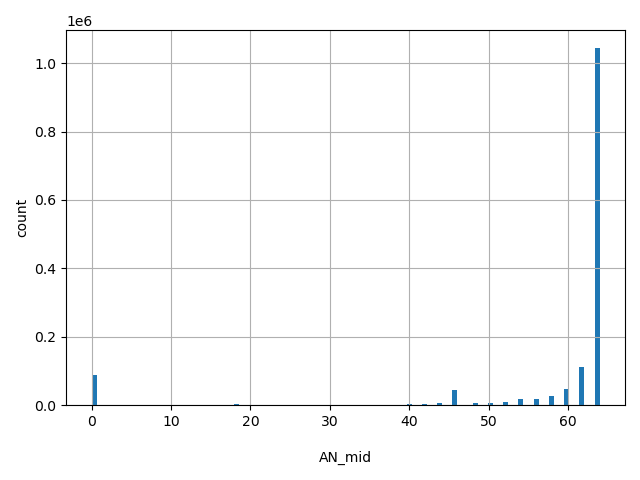 |
[0, 64] |
| AN_nfe |
int |
AN_nfe
|
Total number of Non-Finnish European alleles genotyped (biallelic sites only).
|
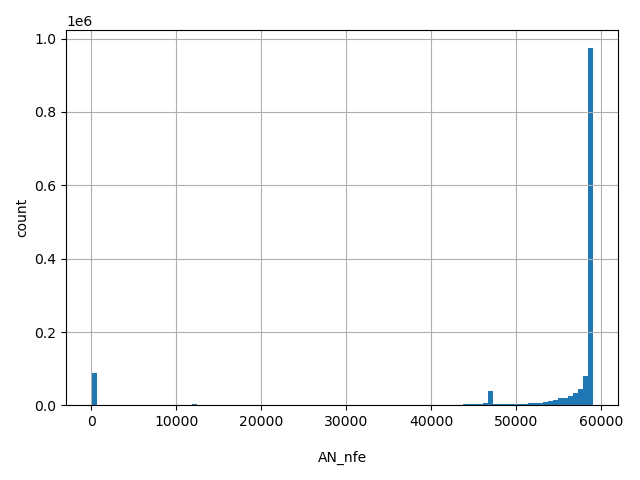 |
[0, 5.91e+04] |
| AN_sas |
int |
AN_sas
|
Total number of South Asian alleles genotyped (biallelic sites only).
|
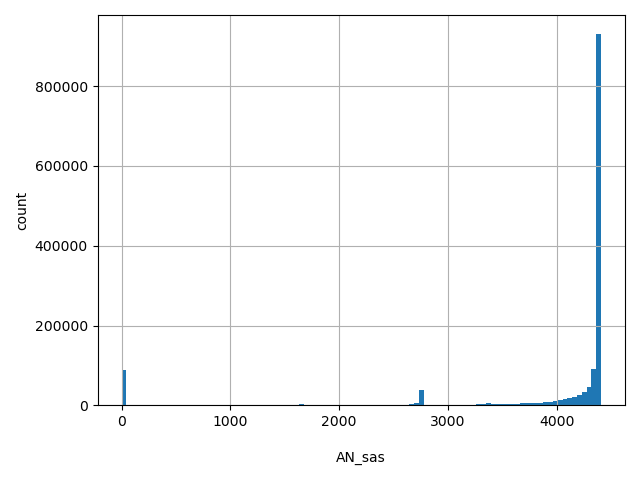 |
[0, 4.41e+03] |
| AC_afr |
int |
AC_afr
|
Number of non-reference African-American alleles observed (biallelic sites only).
|
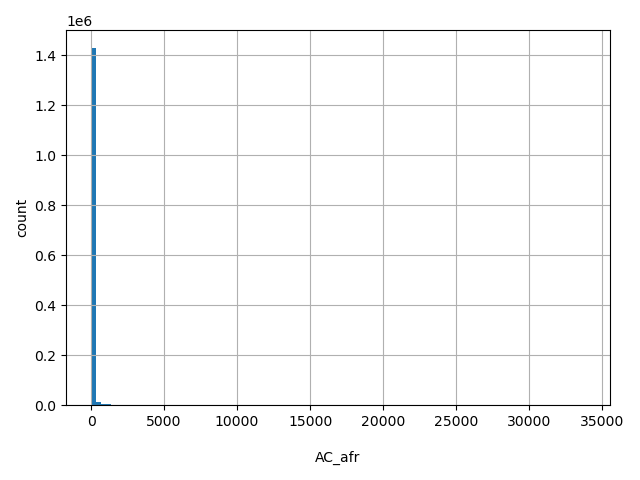 |
[0, 3.38e+04] |
| AC_amr |
int |
AC_amr
|
Number of non-reference Latino alleles observed (biallelic sites only).
|
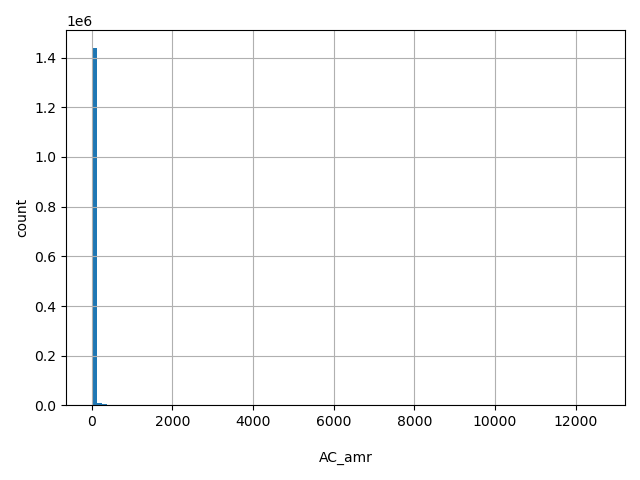 |
[0, 1.26e+04] |
| AC_asj |
int |
AC_asj
|
Number of non-reference Ashkenazi Jewish alleles observed (biallelic sites only).
|
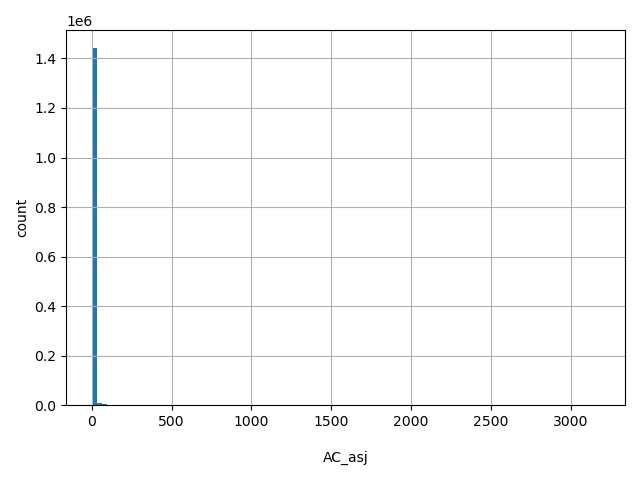 |
[0, 3.18e+03] |
| AC_eas |
int |
AC_eas
|
Number of non-reference East Asian alleles observed (biallelic sites only).
|
 |
[0, 4.05e+03] |
| AC_fin |
int |
AC_fin
|
Number of non-reference Finnish alleles observed (biallelic sites only).
|
 |
[0, 6.48e+03] |
| AC_mid |
int |
AC_mid
|
Number of non-reference Middle Eastern alleles observed (biallelic sites only).
|
 |
[0, 64] |
| AC_nfe |
int |
AC_nfe
|
Number of non-reference Non-Finnish European alleles observed (biallelic sites only).
|
 |
[0, 5.91e+04] |
| AC_sas |
int |
AC_sas
|
Number of non-reference South Asian alleles observed (biallelic sites only).
|
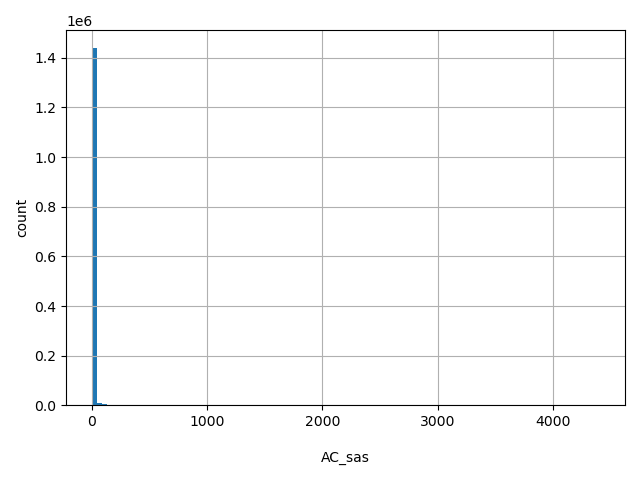 |
[0, 4.41e+03] |
| AF_afr |
float |
AF_afr
|
African-American allele frequency (biallelic sites only).
|
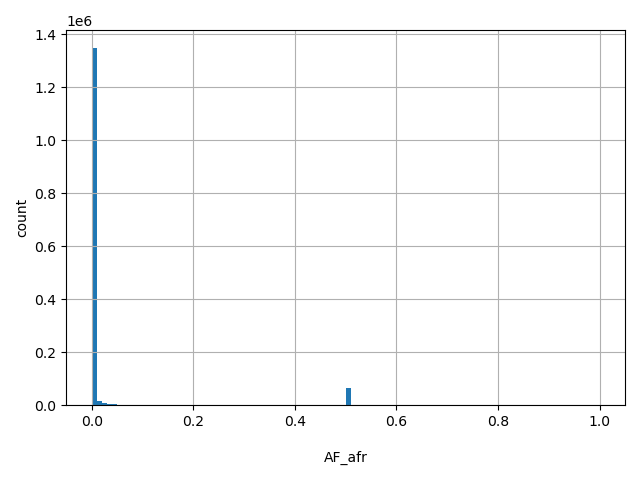 |
[0, 1] |
| AF_amr |
float |
AF_amr
|
Latino allele frequency (biallelic sites only).
|
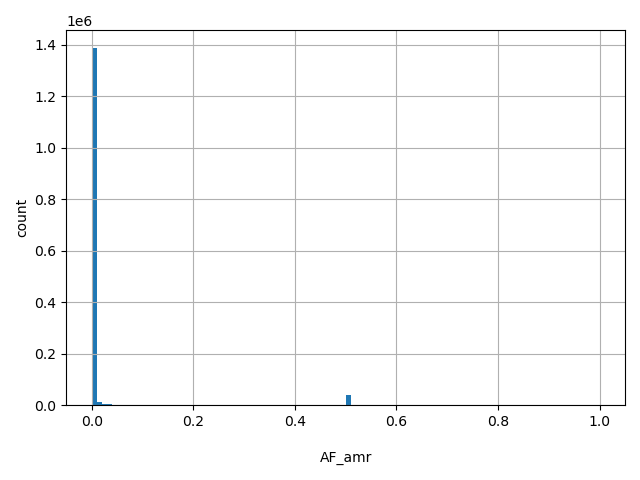 |
[0, 1] |
| AF_asj |
float |
AF_asj
|
Ashkenazi Jewish allele frequency (biallelic sites only).
|
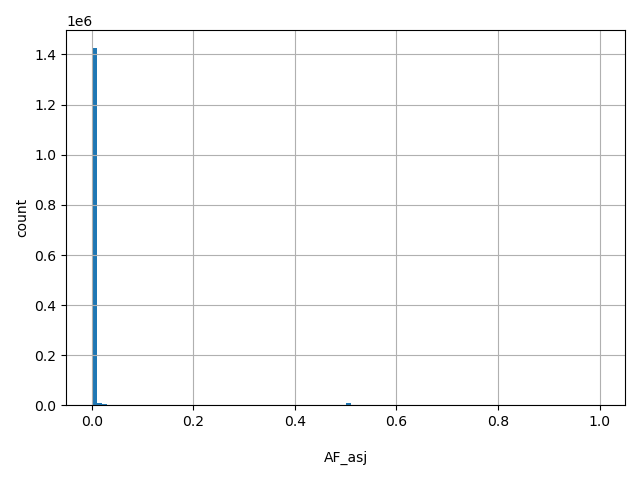 |
[0, 1] |
| AF_eas |
float |
AF_eas
|
East Asian allele frequency (biallelic sites only).
|
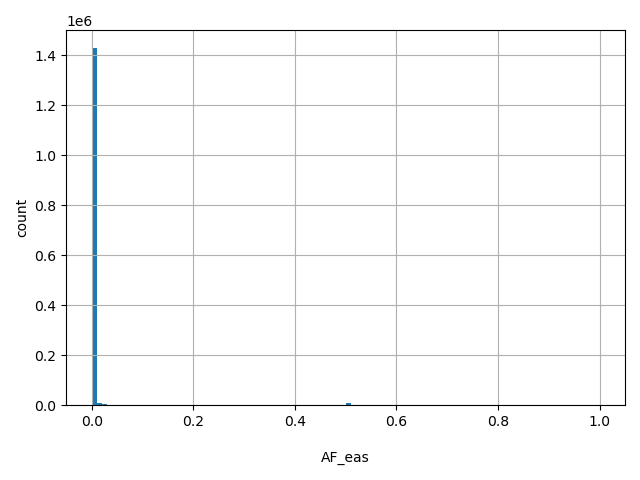 |
[0, 1] |
| AF_fin |
float |
AF_fin
|
Finnish allele frequency (biallelic sites only).
|
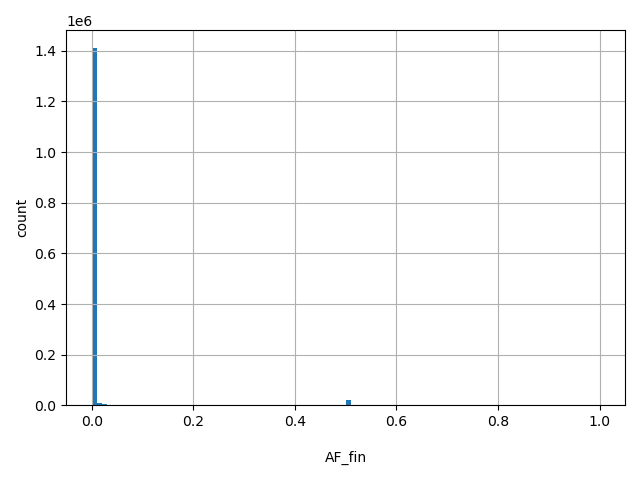 |
[0, 1] |
| AF_mid |
float |
AF_mid
|
Middle Eastern allele frequency (biallelic sites only).
|
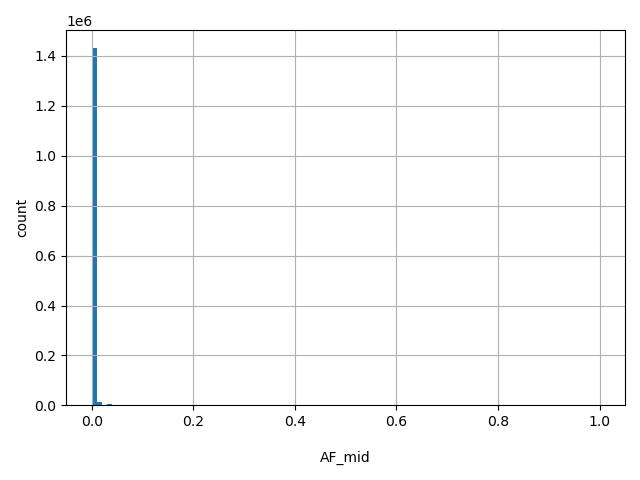 |
[0, 1] |
| AF_nfe |
float |
AF_nfe
|
Non-Finnish European allele frequency (biallelic sites only).
|
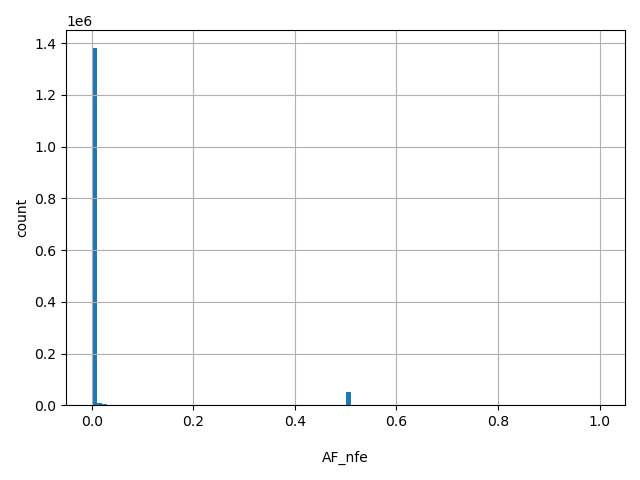 |
[0, 1] |
| AF_sas |
float |
AF_sas
|
South Asian allele frequency (biallelic sites only).
|
 |
[0, 1] |
×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×






























