| ID |
Type |
Default annotation |
Description |
Histogram |
Range |
| deletion_duplication |
str |
deletion_duplication
|
duplication or deletion
|
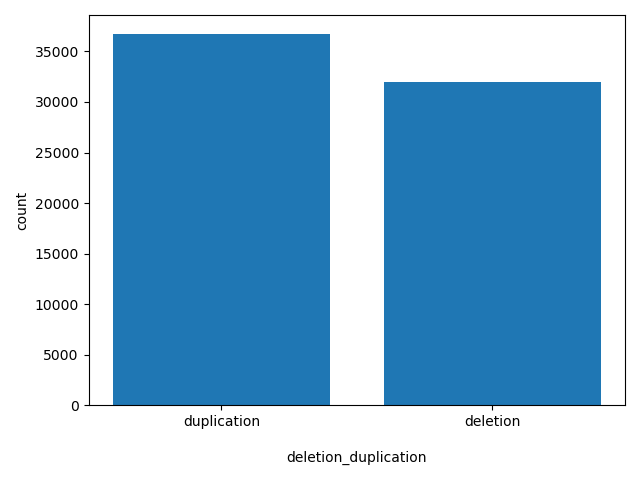 |
duplication, deletion |
| cnv_name |
str |
cnv_name
|
Handy name to refer to the CNV.
|
No histogram: Empty histogram for cnv_name in a region: Too many unique values 101 for categorical histogram. |
NO DOMAIN |
| SVLEN |
int |
SVLEN
|
Length of CNV
|
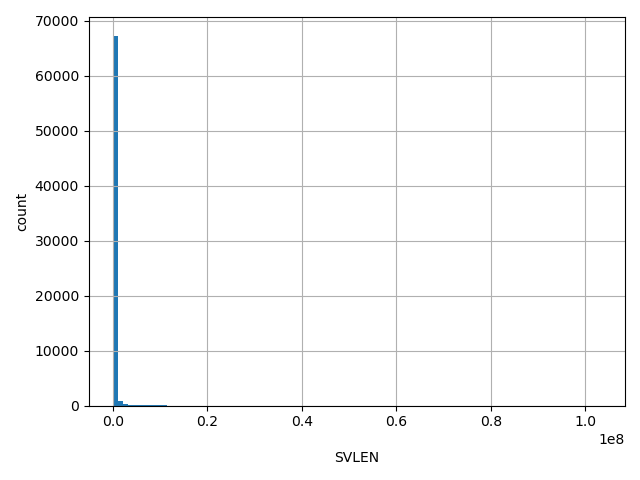 |
[276, 1.03e+08] |
| SC |
int |
SC
|
Number of releasable samples with a variant at this site
|
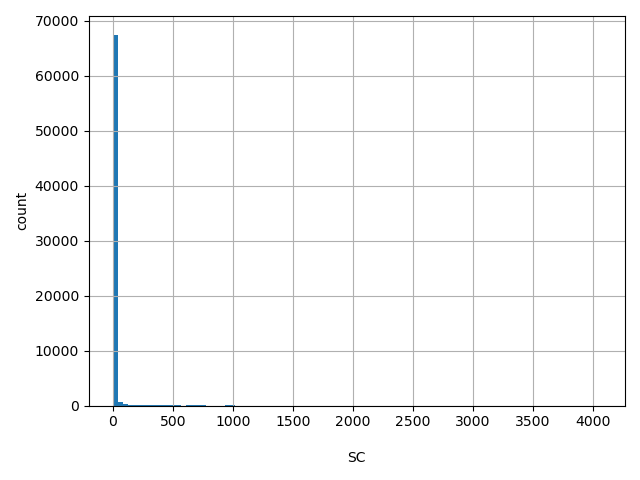 |
[1, 4.07e+03] |
| SN |
int |
SN
|
Total number of releasable samples considered at this site
|
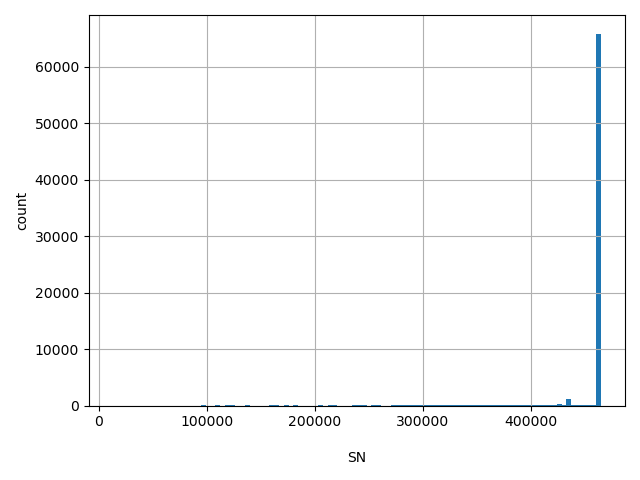 |
[1.32e+04, 4.64e+05] |
| SF |
float |
SF
|
Proportion of releasable samples (site frequency) with a variant at this site
|
 |
[2.15e-06, 0.00876] |
| SC_afr |
int |
SC_afr
|
Number of releasable AFR samples with a variant at this site
|
 |
[0, 1.28e+03] |
| SC_amr |
int |
SC_amr
|
Number of releasable AMR samples with a variant at this site
|
 |
[0, 258] |
| SC_asj |
int |
SC_asj
|
Number of releasable ASJ samples with a variant at this site
|
 |
[0, 551] |
| SC_eas |
int |
SC_eas
|
Number of releasable EAS samples with a variant at this site
|
 |
[0, 615] |
| SC_fin |
int |
SC_fin
|
Number of releasable FIN samples with a variant at this site
|
 |
[0, 725] |
| SC_mid |
int |
SC_mid
|
Number of releasable MID samples with a variant at this site
|
 |
[0, 79] |
| SC_nfe |
int |
SC_nfe
|
Number of releasable NFE samples with a variant at this site
|
 |
[0, 3.49e+03] |
| SC_sas |
int |
SC_sas
|
Number of releasable SAS samples with a variant at this site
|
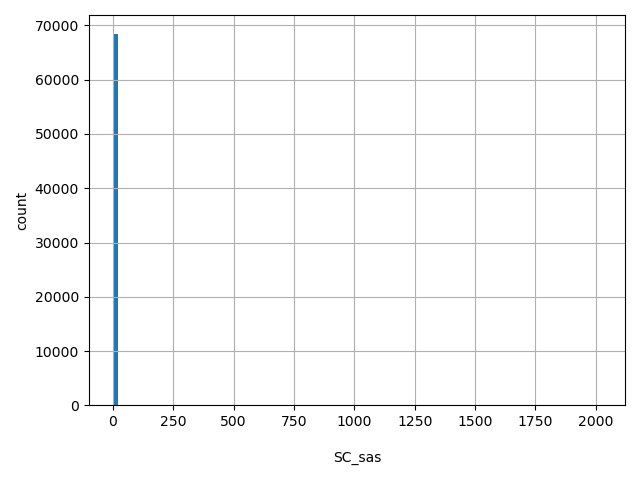 |
[0, 2.02e+03] |
| SN_afr |
int |
SN_afr
|
Total number of releasable AFR samples considered at this site
|
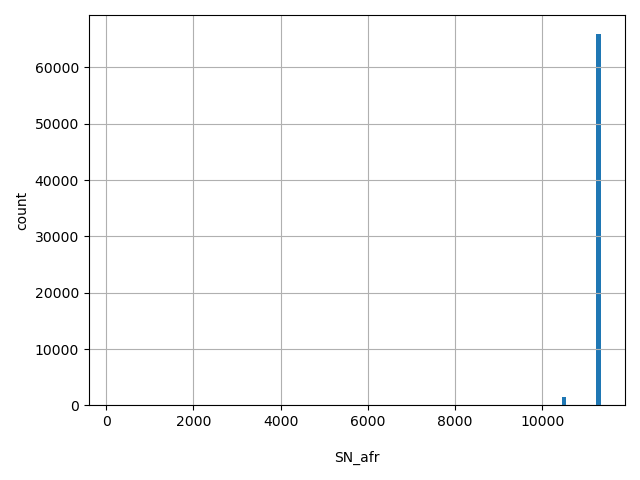 |
[157, 1.13e+04] |
| SN_amr |
int |
SN_amr
|
Total number of releasable AMR samples considered at this site
|
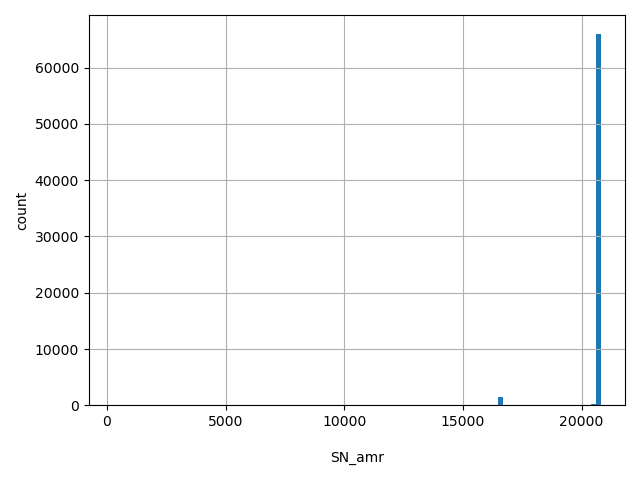 |
[253, 2.08e+04] |
| SN_asj |
int |
SN_asj
|
Total number of releasable ASJ samples considered at this site
|
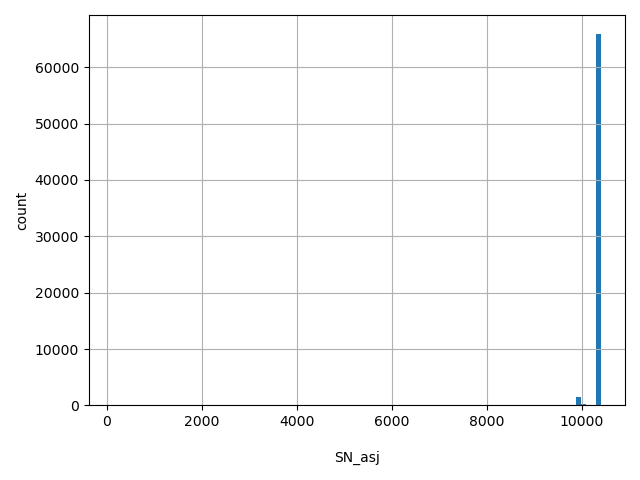 |
[128, 1.04e+04] |
| SN_eas |
int |
SN_eas
|
Total number of releasable EAS samples considered at this site
|
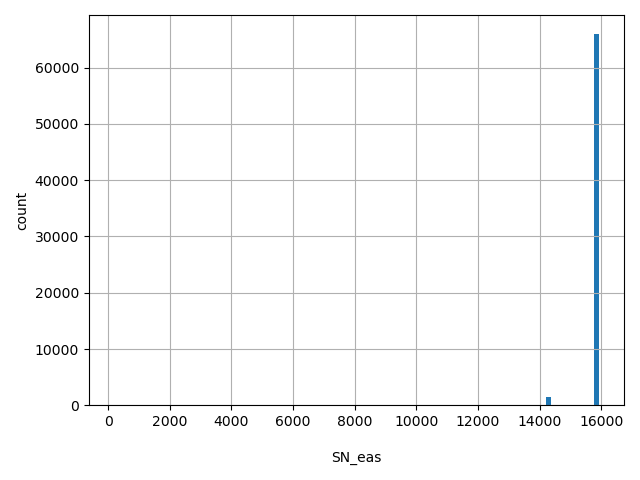 |
[159, 1.59e+04] |
| SN_fin |
int |
SN_fin
|
Total number of releasable FIN samples considered at this site
|
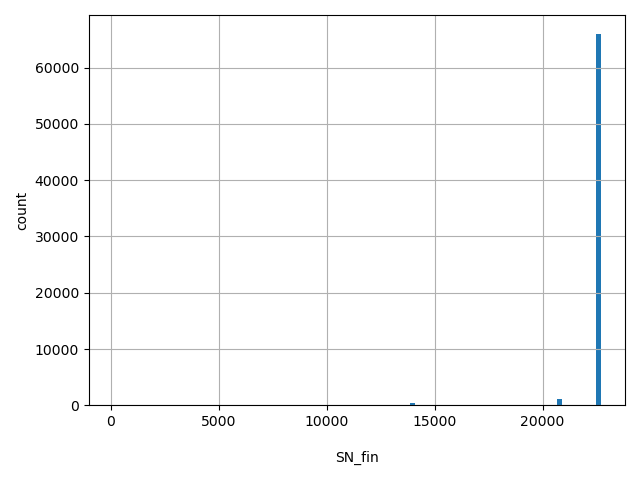 |
[106, 2.27e+04] |
| SN_mid |
int |
SN_mid
|
Total number of releasable MID samples considered at this site
|
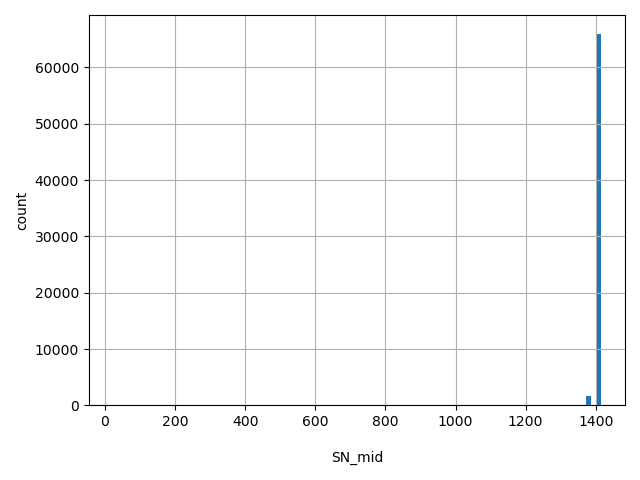 |
[23, 1.41e+03] |
| SN_nfe |
int |
SN_nfe
|
Total number of releasable NFE samples considered at this site
|
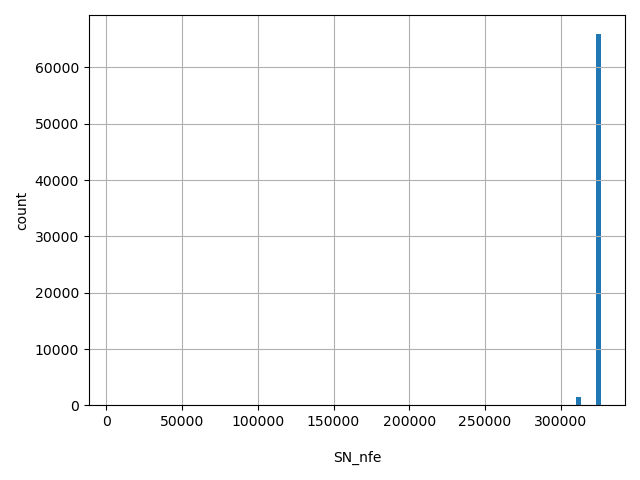 |
[4.44e+03, 3.26e+05] |
| SN_sas |
int |
SN_sas
|
Total number of releasable SAS samples considered at this site
|
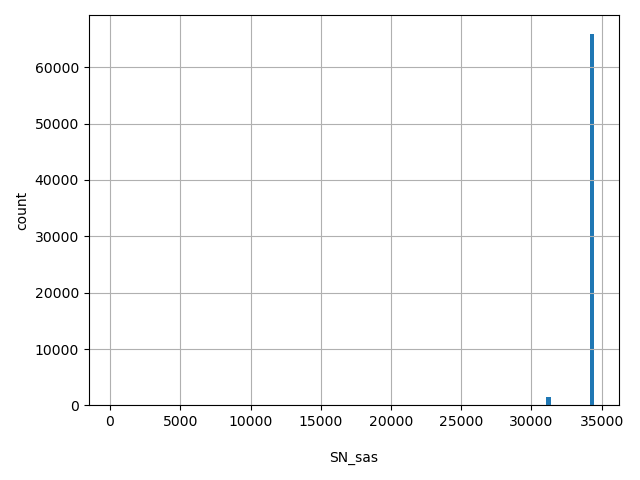 |
[191, 3.45e+04] |
| SF_afr |
float |
SF_afr
|
Proportion of releasable AFR samples with a variant at this site
|
 |
[0, 0] |
| SF_amr |
float |
SF_amr
|
Proportion of releasable AMR samples with a variant at this site
|
 |
[0, 0] |
| SF_asj |
float |
SF_asj
|
Proportion of releasable ASJ samples with a variant at this site
|
 |
[0, 0] |
| SF_eas |
float |
SF_eas
|
Proportion of releasable EAS samples with a variant at this site
|
 |
[0, 0] |
| SF_fin |
float |
SF_fin
|
Proportion of releasable FIN samples with a variant at this site
|
 |
[0, 0] |
| SF_mid |
float |
SF_mid
|
Proportion of releasable MID samples with a variant at this site
|
 |
[0, 0] |
| SF_nfe |
float |
SF_nfe
|
Proportion of releasable NFE samples with a variant at this site
|
 |
[0, 0] |
| SF_sas |
float |
SF_sas
|
Proportion of releasable SAS samples with a variant at this site
|
 |
[0, 0] |
×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×

×






























