Resource
| Id | hg38/cnv_collections/dbVar |
|---|---|
| Type | cnv_collection |
| Version | 0 |
| Summary |
dbVar CNV collection
|
| Description |
dbVar is a database of genomic structural variations, including deletions, duplications, inversions, and copy number variations (CNVs), collected from human and model organism studies. It provides curated data on variant regions (generalized variation locations) and variant calls (specific observed instances), aiding in understanding genetic diversity and disease associations. Downloaded on 12/16/2024 from https://ftp.ncbi.nlm.nih.gov/pub/dbVar/data/Homo_sapiens/by_study/csv/variants_for_nstd186.csv.gz DataPrep.py converts raw file to GRR format. 'Start' and 'Inner Start" columns are consolidated to "pos_beg"; "End" and "Inner End" columns are consolidated to "pos_end". |
| Labels |
Scores (2)
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| deletion_duplication | str |
deletion_duplication |
duplication or deletion
|
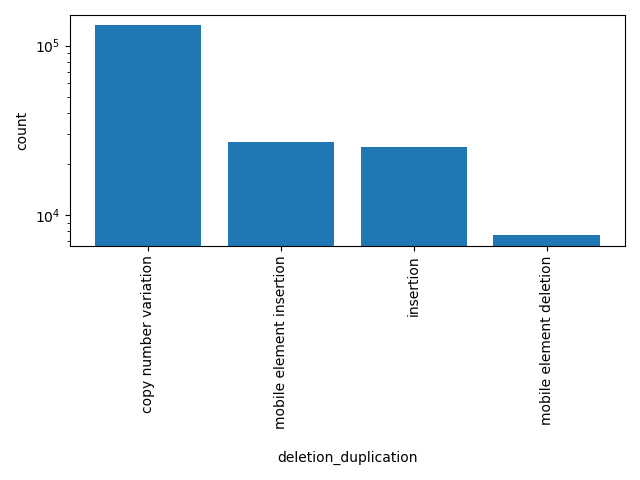 |
copy number variation, mobile element insertion, insertion, mobile element deletion |
| cnv_name | str |
cnv_name |
Handy name to refer to the CNV.
|
No histogram: Empty histogram for cnv_name in a region: Too many unique values 101 for categorical histogram. |
NO DOMAIN |
Files
| Filename | Size | md5 |
|---|---|---|
| DataPrep.py | 1.17 KB | 6486f4a94df34fb5acd40481075dec2d |
| genomic_resource.yaml | 1.57 KB | 2ad2ba4e4577226b257b06eacc2b22f8 |
| prepped_variants_for_nstd186.csv | 19.94 MB | 923ff1361a06190174962fd401c11dbe |
| prepped_variants_for_nstd186.csv.dvc | 116.0 B | 21e7c8d643ec6c150c035335147dc2e9 |
| prepped_variants_for_nstd186.tsv.gz | 2.23 MB | 5393e43f9cc6e1dbaf7a1a1598c1ce23 |
| variants_for_nstd186.csv | 21.23 MB | fce0401ee70a29a0dcaebe4cb535e3ca |
| variants_for_nstd186.csv.dvc | 108.0 B | e585d43ed183ee8d4bec8869c43afc26 |
| statistics/ |
