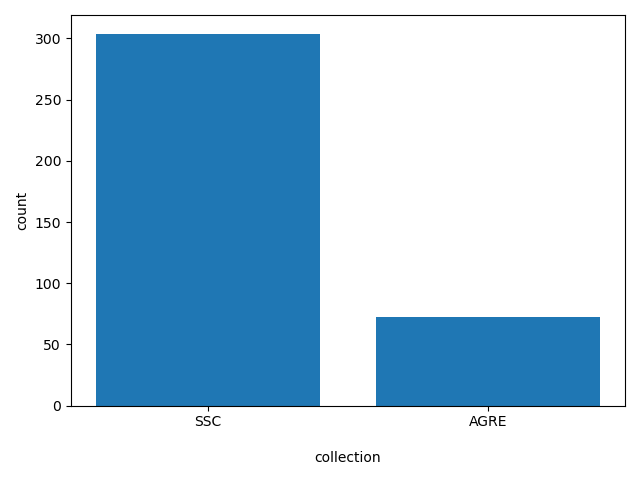
collection
SSC or AGRE
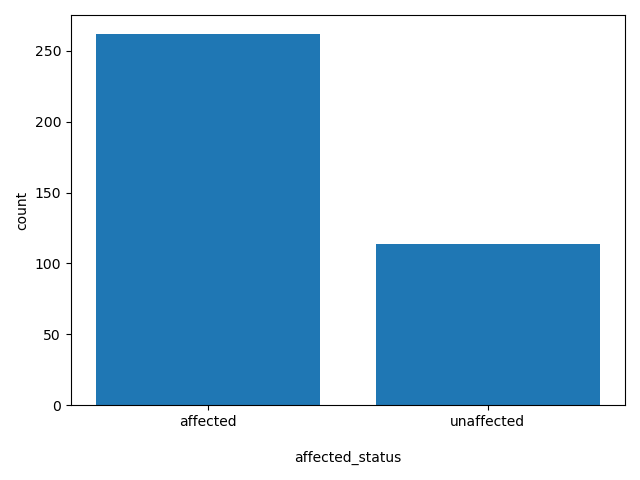
affected_status
Shows if the child that has the de novo is affected or unaffected.
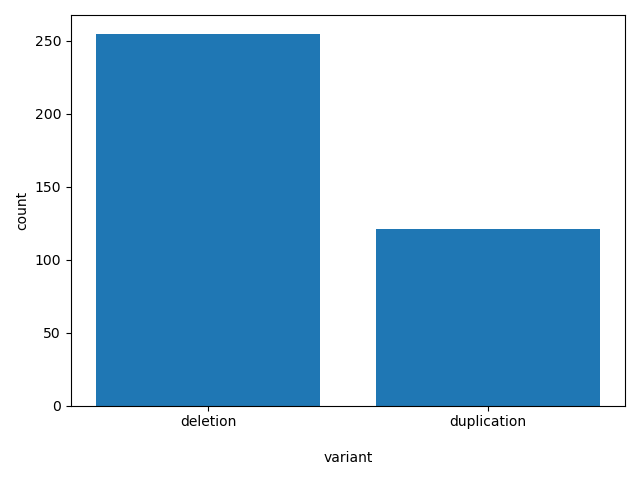
variant
duplication or deletion
| Id | hg38/cnv_collections/Iossifov_Lab_SSC_AGRE_2021_new |
|---|---|
| Type | cnv_collection |
| Version | 0 |
| Summary |
De novo CNVs from SSC and AGRE WGS
|
| Description |
The Iossifov Lab AGRE-SSC CNV collection is a curated table of de novo copy-number variants (CNVs) assembled from the whole-genome sequencing study by Yoon et al. (2021), which analyzed WGS data from simplex families in the Simons Simplex Collection (SSC) and multiplex families in the Autism Genetic Resource Exchange (AGRE). Each record is a CNV interval with its class (deletion or duplication), size, and basic overlap metadata (e.g., gene count/genes). Supplementary Table 4 downloaded on: 02/20/26 |
| Labels |
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| collection | str |
collection |
SSC or AGRE
|
 |
SSC, AGRE |
| affected_status | str |
affected_status |
Shows if the child that has the de novo is affected or unaffected.
|
 |
affected, unaffected |
| variant | str |
variant |
duplication or deletion
|
 |
deletion, duplication |
| Filename | Size | md5 |
|---|---|---|
| 42003_2021_2533_MOESM6_ESM.xlsx | 54.26 KB | 92e09df3ea3da4d68913b3f56aa0e0c1 |
| Iossifov_Lab_SSC_AGRE_2021.tsv | 17.49 KB | 385b3ed82221d0af6a708f8cba448c7a |
| dataprep.py | 1.37 KB | b6657d0cf0e4924448ed7ba35f027999 |
| genomic_resource.yaml | 1.33 KB | 4d063f52c1f771e72c011436cddb7832 |
| statistics/ |