| Id | gene_properties/gene_sets/geneimprint |
|---|---|
| Type | gene_set_collection |
| Version | 0 |
| Summary |
geneimprint human gene sets
|
| Description |
geneimprint provides a curated catalog of imprinted genes and related imprinting information across species. Genomic imprinting is an epigenetic form of gene regulation in which only one parental copy of a gene is preferentially expressed, depending on whether it was inherited from the mother or the father. Because imprinted genes can functionally depend on a single active parental allele, changes affecting that allele or its epigenetic regulation can have important developmental and disease-related consequences. GeneImprint organizes imprinting-related gene information by species, gene, imprinting status, chromosomal location, and expressed parental allele, providing a practical reference for identifying and analyzing genes with known or predicted imprinting behavior. This gene set collection is generated by dataprep.py, which reads the geneimprint genes-by-species table from the website, extracts the human gene entries, and writes them in GMT format. The resulting file contains gene sets grouped by imprinting status, expressed allele, combined status and expressed allele, and an all_genes set containing all human genes in the table. Downloaded on: 05/04/26 https://www.geneimprint.com/site/genes-by-species.Homo+sapiens |
| Labels |
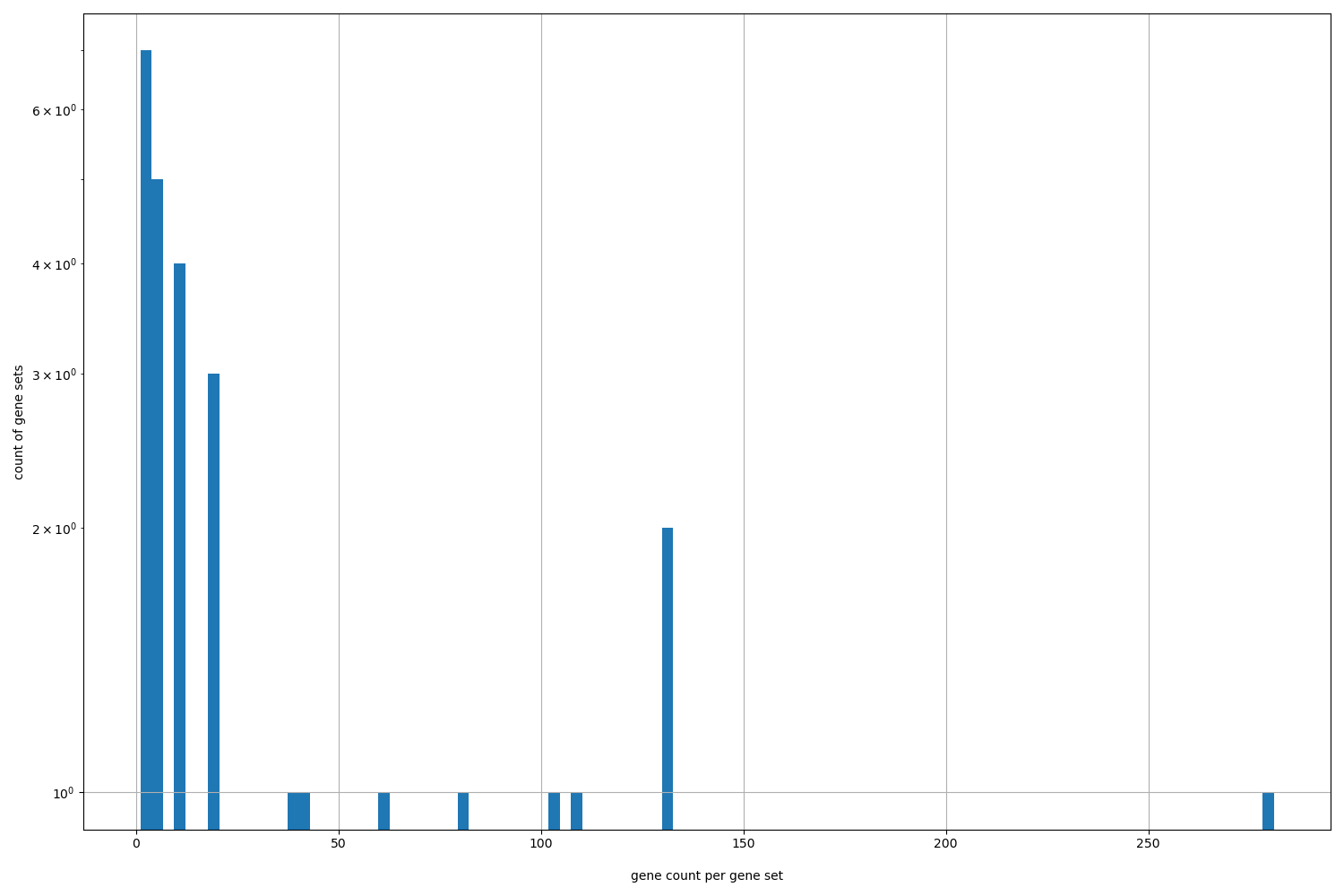
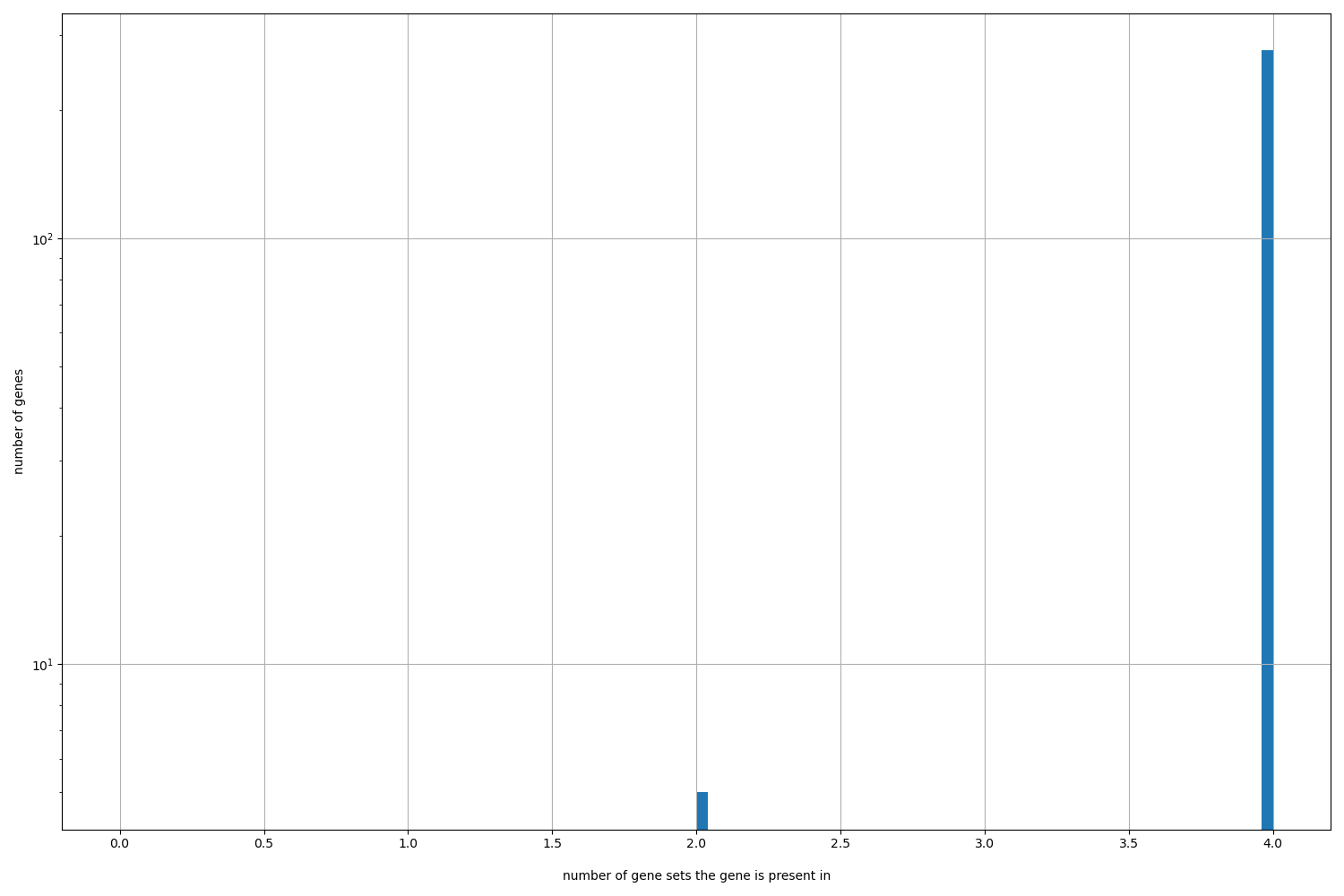
Number of gene sets: 28
Number of unique genes: 281


| Gene Set | Gene Count | Description |
|---|---|---|
| all_genes | 281 | all_genes |
| STATUS_imprinted | 131 | STATUS_imprinted |
| EXPRESSED_ALLELE_paternal | 130 | EXPRESSED_ALLELE_paternal |
| EXPRESSED_ALLELE_maternal | 109 | EXPRESSED_ALLELE_maternal |
| STATUS_predicted | 104 | STATUS_predicted |
| STATUS_imprinted_EXPRESSED_ALLELE_paternal | 80 | STATUS_imprinted_EXPRESSED_ALLELE_paternal |
| STATUS_predicted_EXPRESSED_ALLELE_maternal | 62 | STATUS_predicted_EXPRESSED_ALLELE_maternal |
| STATUS_predicted_EXPRESSED_ALLELE_paternal | 42 | STATUS_predicted_EXPRESSED_ALLELE_paternal |
| STATUS_imprinted_EXPRESSED_ALLELE_maternal | 40 | STATUS_imprinted_EXPRESSED_ALLELE_maternal |
| EXPRESSED_ALLELE_biallelic | 20 | EXPRESSED_ALLELE_biallelic |
| STATUS_not_imprinted | 18 | STATUS_not_imprinted |
| STATUS_not_imprinted_EXPRESSED_ALLELE_biallelic | 18 | STATUS_not_imprinted_EXPRESSED_ALLELE_biallelic |
| EXPRESSED_ALLELE_unknown | 11 | EXPRESSED_ALLELE_unknown |
| STATUS_unknown | 11 | STATUS_unknown |
| STATUS_unknown_EXPRESSED_ALLELE_unknown | 11 | STATUS_unknown_EXPRESSED_ALLELE_unknown |
| STATUS_conflicting_data | 10 | STATUS_conflicting_data |
| STATUS_conflicting_data_EXPRESSED_ALLELE_paternal | 5 | STATUS_conflicting_data_EXPRESSED_ALLELE_paternal |
| STATUS_provisional_data | 5 | STATUS_provisional_data |
| EXPRESSED_ALLELE_isoform_dependent | 4 | EXPRESSED_ALLELE_isoform_dependent |
| STATUS_imprinted_EXPRESSED_ALLELE_isoform_dependent | 4 | STATUS_imprinted_EXPRESSED_ALLELE_isoform_dependent |
| STATUS_provisional_data_EXPRESSED_ALLELE_maternal | 4 | STATUS_provisional_data_EXPRESSED_ALLELE_maternal |
| STATUS_conflicting_data_EXPRESSED_ALLELE_maternal | 3 | STATUS_conflicting_data_EXPRESSED_ALLELE_maternal |
| EXPRESSED_ALLELE_random | 2 | EXPRESSED_ALLELE_random |
| STATUS_conflicting_data_EXPRESSED_ALLELE_biallelic | 2 | STATUS_conflicting_data_EXPRESSED_ALLELE_biallelic |
| STATUS_imprinted_EXPRESSED_ALLELE_random | 2 | STATUS_imprinted_EXPRESSED_ALLELE_random |
| STATUS_tissue_dependent | 2 | STATUS_tissue_dependent |
| STATUS_tissue_dependent_EXPRESSED_ALLELE_paternal | 2 | STATUS_tissue_dependent_EXPRESSED_ALLELE_paternal |
| STATUS_provisional_data_EXPRESSED_ALLELE_paternal | 1 | STATUS_provisional_data_EXPRESSED_ALLELE_paternal |
Format: gmt
| Filename | Size | md5 |
|---|---|---|
| dataprep.py | 2.39 KB | 3809324a146e8057812f996912fd81c8 |
| geneimprint_human.gmt | 9.04 KB | 6f7af7766297ccc3c160dafb693727ee |
| geneimprint_table.csv | 16.63 KB | f600d01266956ab7ebacc05bf8a7de5f |
| genomic_resource.yaml | 1.66 KB | bbfd23c4b2bb8cde438a55ba02b00eee |
| statistics/ |