| Id | gene_properties/gene_scores/RVIS |
|---|---|
| Type | gene_score |
| Version | 0 |
| Summary |
Residual Variation Intolerance Score
|
| Description |
The Residual Variation Intolerance Score (RVIS) is a gene-level metric that quantifies how tolerant a gene is to functional genetic variation in the general population. RVIS measures the deviation between the observed and expected number of common functional variants per gene, after accounting for overall mutational burden. Genes with lower RVIS values are more variation-intolerant and often linked to severe Mendelian disease, whereas higher values indicate greater tolerance and reduced disease likelihood. Downloaded on: 10/23/25 Processing details: dataPrep.py renames first column of Supplementary Dataset S2 from "HGNC gene" to "gene", and saves as a csv file. |
| Labels |
| ID | Type | Description | Histograms | Range |
|---|---|---|---|---|
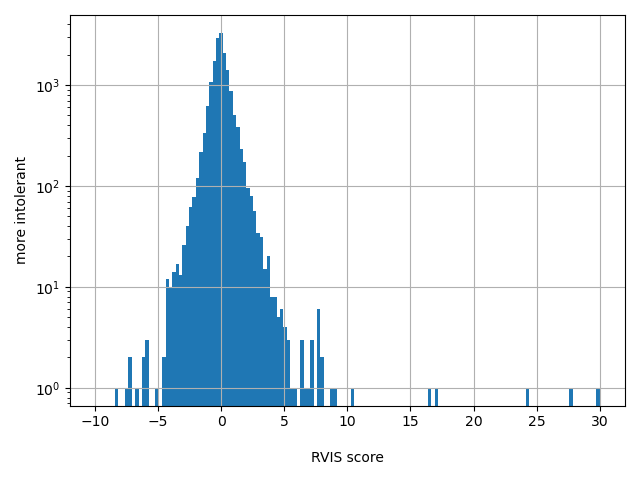
| RVIS score |
Intolerance of a gene to genetic variants
Small values desc: more intolerant
Large values desc: less intolerant
|

|
[-8.29, 29.8] | |
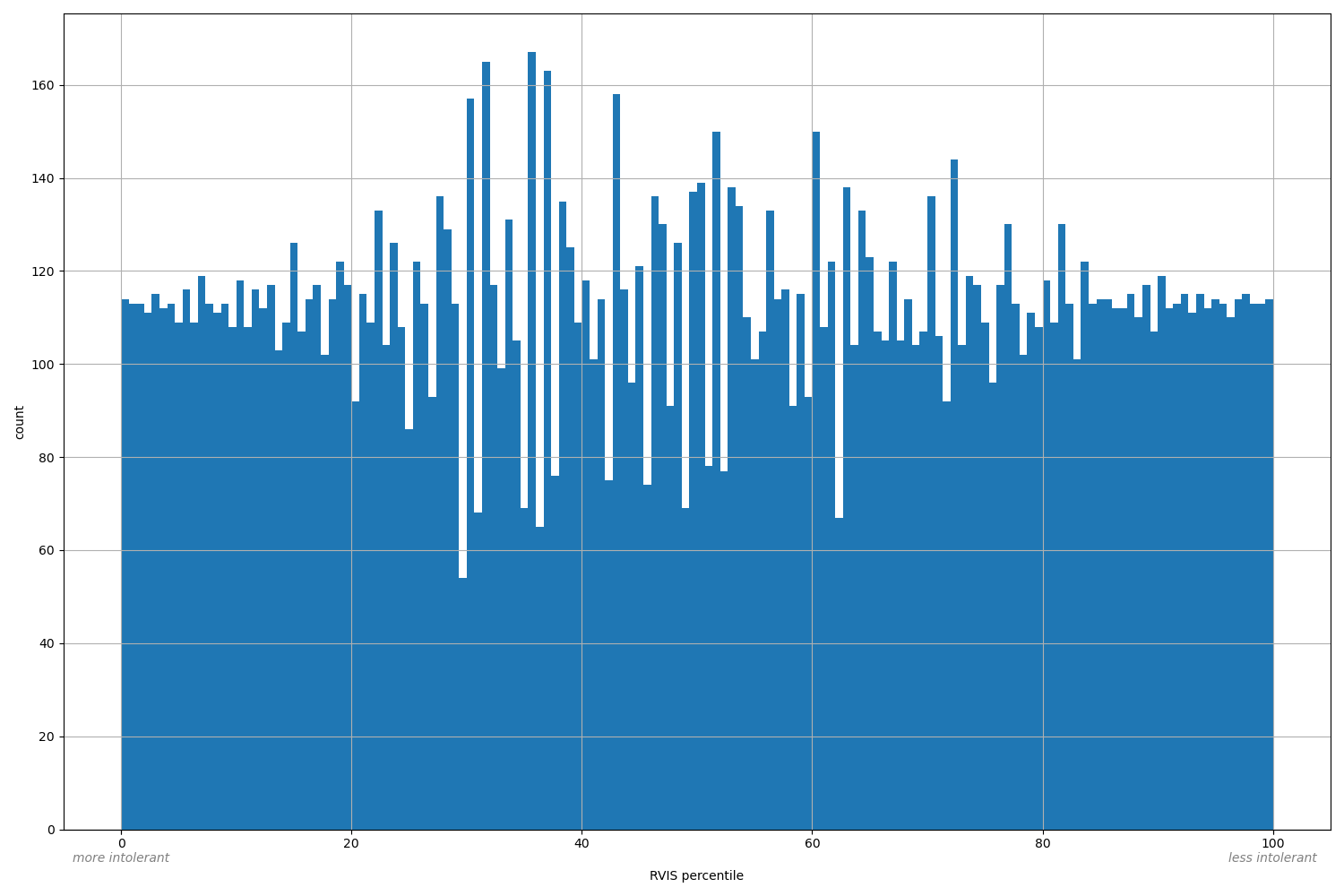
| RVIS percentile |
Gene percentile after sorting by RVIS intolerance score
Small values desc: more intolerant
Large values desc: less intolerant
|

|
[0.0059, 100] |
| Filename | Size | md5 |
|---|---|---|
| RVIS.csv | 712.81 KB | cdba62f48f47c4476c39973c9bca97f4 |
| dataprep.py | 148.0 B | e3e903cb5f7766a2a0f0a8c95378f988 |
| genomic_resource.yaml | 1.96 KB | 9f93ef2903a9d697bf52cc42c2c99468 |
| pgen.1003709.s002.xlsx | 750.44 KB | d333c4dd1d2f4c928fd4c7ccc6f29a25 |
| pgen.1003709.s002.xlsx.dvc | 104.0 B | 33bb739167f3e9995008456058b958a5 |
| statistics/ |